File:3D-fluorescence imaging for high throughput analysis of microbial eukaryotes.jpg
From Wikimedia Commons, the free media repository
Jump to navigation
Jump to search

Size of this preview: 800 × 515 pixels. Other resolutions: 320 × 206 pixels | 640 × 412 pixels | 1,024 × 660 pixels | 1,280 × 825 pixels | 2,560 × 1,650 pixels | 5,098 × 3,285 pixels.
Original file (5,098 × 3,285 pixels, file size: 1.39 MB, MIME type: image/jpeg)
File information
Structured data
Captions
Captions
3D-fluorescence imaging for high throughput analysis of microbial eukaryotes
Summary
[edit]| Description3D-fluorescence imaging for high throughput analysis of microbial eukaryotes.jpg |
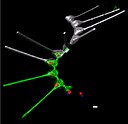
English: Environmental high-content fluorescence microscopy (e-HCFM): automated, 3D, and multichannel imaging for aquatic micro-eukaryotes. (a) e-HCFM workflow applied to Tara Oceans samples: (1) 72 nano-plankton (size range 5–20 μm) samples collected during the Tara Oceans expedition (Pesant et al., 2015) were fixed in paraformaldehyde-glutaraldehyde buffer onboard and kept at 4°C for up to several years; (2) Samples were mounted in optical multiwell plates. Then, a 4-steps preparation allowed to stain all eukaryotic cells; (3) A commercial confocal laser scanning microscope was used to automatically image samples (40X NA1.1 water lens; 5 channels) generating 2.5 Tb of raw data (acquisition details in Figure 1—source data 1); (4) In total, 336,655 objects were processed for individual extraction of 480 descriptors (3D biovolumes, intensity distribution, shape descriptors and texture features, details in Figure 1—source data 2), and the reconstruction of various images for visual inspection (c); (5) A training set based on 18,103 manually curated images (5.4% of the dataset) classified into 155 categories, was used for automated recognition (Random Forest). (b) Examples of e-HCFM 3D-images and movies from a wide phylogenetic diversity of planktonic eukaryotes (see also Figure 1—figure supplements 1 and 2). Left panel: a chain of diatoms (Asterionellopsis sp., Heterokonta) (Figure 1—video 1); right panel, top left to bottom right: radiolarian (Rhizaria), ciliate (Alveolata), amoeba (Amoebozoa), choanoflagellate (Opisthokonta), dinoflagellate (Alveolata), coccolithophore (Haptophyta). Key cellular features are labelled with various dyes: DNA/nuclei (blue, Hoechst33342); (intra)cellular membranes (green, DiOC6(3)); cell covers and extensions (cyan, PLL-AF546, a home-made conjugation between α-poly-L-lysine (PLL) and Alexa Fluor 546 (AF546)); chloroplasts (red, chlorophyll autofluorescence). Scale bar 5 µm. (c) The confocal microscope is automated for acquiring multicolor Z-stacks over a mosaic of positions for each sample. |
|||
| Date | ||||
| Source | doi:10.7554/eLife.26066.003 | |||
| Author | Sebastien Colin, Luis Pedro Coelho, Shinichi Sunagawa, Chris Bowler, Eric Karsenti, Peer Bork, Rainer Pepperkok, Colomban de Vargas | |||
| Permission (Reusing this file) |
|
|||
| Other versions |
|
Licensing
[edit]This file is licensed under the Creative Commons Attribution 4.0 International license.
- You are free:
- to share – to copy, distribute and transmit the work
- to remix – to adapt the work
- Under the following conditions:
- attribution – You must give appropriate credit, provide a link to the license, and indicate if changes were made. You may do so in any reasonable manner, but not in any way that suggests the licensor endorses you or your use.
File history
Click on a date/time to view the file as it appeared at that time.
| Date/Time | Thumbnail | Dimensions | User | Comment | |
|---|---|---|---|---|---|
| current | 22:11, 19 October 2020 |  | 5,098 × 3,285 (1.39 MB) | Remitamine (talk | contribs) | Higher resolution version |
| 07:57, 5 October 2020 |  | 1,500 × 966 (256 KB) | Epipelagic (talk | contribs) | Uploaded a work by Sebastien Colin, Luis Pedro Coelho, Shinichi Sunagawa, Chris Bowler, Eric Karsenti, Peer Bork, Rainer Pepperkok, Colomban de Vargas from [https://elifesciences.org/articles/26066] {{doi|https://doi.org/10.7554/eLife.26066.003}} with UploadWizard |
You cannot overwrite this file.
File usage on Commons
The following page uses this file:
File usage on other wikis
The following other wikis use this file:
- Usage on en.wikipedia.org